Cotton is the world’s most important renewable textile fiber and a significant oilseed crop [9]. Cotton fiber is widely used for apparel, home furnishings, and other diverse industrial products. World consumption of cotton fiber is ~115-million bales or ~27-million metric tons per year (National Cotton Council, http://www.cotton.org/, 2008). Improving cotton fiber yield and quality will provide a sustainable alternative to petroleum-based synthetic fibers. Efforts are also ongoing to modify cottonseed for food and feed [7, 10], which could enhance nutrition and livelihoods of millions of people in food-challenged economies. The current predominance of transgenic varieties that require less chemical application while increasing yield, together with research efforts to increase the abiotic and biotic stress resistance, predict that economically viable cotton crops can be grown in a sustainable manner in many warm global locations that also constitute major population centers in the developing world. Fully understanding the control of fiber development is a major goal to support future improvements in cotton fiber yield and quality as a cornerstone of this global vision.
The genus Gossypium includes ~45 diploid (2n = 2x = 26) and five tetraploid (2n = 4x = 52) species. Diploid species fall into eight genomic groups (A-G & K). The African clade (A, B, E, and F genomes [8]) occurs naturally in Africa and Asia, while the D-genome clade is indigenous to the Americas. A third diploid clade (C, G, and K genomes) is found in Australia. Allotetraploids arose in the New World from interspecific hybridization between an A-genome-like African species and a D-genome-like American species, which occurred 1-2 million years ago (Mya) [11] (Figure 1). The closest relatives of the original tetraploid progenitors are the A-genome species Gossypium herbaceum L. (A1) andGossypium arboreum L. (A2) and the D-genome species Gossypium raimondii (D5) Ulbrich. Two of five allotetraploid species, Gossypium hirsutum L. and Gossypium barbadense L. are domesticated and cultivated. Interestingly, the A-genome species produce spinnable fiber and are cultivated on a limited scale, whereas the D-genome fiber is rudimentary and not useful [2]. The fiber in allotetraploids is much longer and stronger, suggesting activation or silencing of homoeologous fiber-related genes by genetic and epigenetic mechanisms [1, 3, 6, 11], leading to hybrid vigor [4].
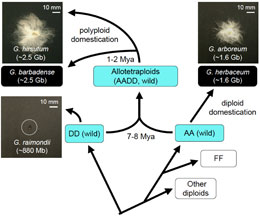 Figure 1. Formation of allotetraploid cotton. Extant diploid progenitors diverge 7-8 millions years ago (Mya), and allotetraploidization occurred naturally 1-2 Mya between a fiber-producing AA-genome extant species and a fiber-poor DD-genome extant species, generating AADD allotetraploid species. Superior fiber yield and quality have been selected in allotetraploid cotton species, as well as the domesticated diploid G. arboreum. The genome sizes in parentheses are cited from [5]. Mbp: mega base pairs (106); Gbp: giga base pairs (109).
Figure 1. Formation of allotetraploid cotton. Extant diploid progenitors diverge 7-8 millions years ago (Mya), and allotetraploidization occurred naturally 1-2 Mya between a fiber-producing AA-genome extant species and a fiber-poor DD-genome extant species, generating AADD allotetraploid species. Superior fiber yield and quality have been selected in allotetraploid cotton species, as well as the domesticated diploid G. arboreum. The genome sizes in parentheses are cited from [5]. Mbp: mega base pairs (106); Gbp: giga base pairs (109).
Most production (>97%) in the U.S. is Upland cotton (G. hirsutum L., AD1), and the remainder is Pima cotton (G. barbadense L, AD2) with extra-long fiber that is sought after for high quality textiles. In 2009, over 12.5 million bales (1 bale = 170 kg) of fiber were produced in the US on over 30 thousand farms in 17 states covering >8 million acres, including Texas, California, Mississippi, and North Carolina. Texas alone accounts for 39% of the nation's harvest in 2009. The total value of the annual US raw fiber crop is ~$6 billion, annual values of U.S. cotton sold overseas recently have averaged ~$3 billion (a trade surplus to the US), accounting for ~25% of the total world export market. Total business revenues from related US industries, such as gins, merchants, warehouses, oil and textile mills, account for more than $40 billion. In the US, nearly 450,000 jobs are directly linked to the cotton industry, and business revenue stimulated by the crop is estimated at $120 billion. The targeted introduction of fiber quality traits from G. barbadense into more adaptable and higher yielding G. hirsutum is a major goal for cotton improvement. The research proposed here will support this goal by identifying relevant controlling genes in two species.
References
- Adams KL, Wendel JF: Polyploidy and genome evolution in plants. Curr Opin Plant Biol 8: 135-41 (2005).
- Applequist WL, Cronn R, Wendel JF: Comparative development of fiber in wild and cultivated cotton. Evol Dev 3: 3-17. (2001).
- Chen ZJ: Genetic and epigenetic mechanisms for gene expression and phenotypic variation in plant polyploids. Annu Rev Plant Biol 58: 377-406 (2007).
- Chen ZJ: Molecular mechanisms of polyploidy and hybrid vigor. Trends Plant Sci (10.1016/j.tplants.2009.12.003) (2010).
- Hendrix B, Stewart JM: Estimation of the nuclear DNA content of gossypium species. Ann Bot (Lond) 95: 789-97 (2005).
- Lee JJ, Woodward AW, Chen ZJ: Gene expression changes and early events in cotton fibre development. Ann Bot (Lond) 100: 1391-401 (2007).
- Mao YB, Cai WJ, Wang JW, Hong GJ, Tao XY, Wang LJ, Huang YP, Chen XY: Silencing a cotton bollworm P450 monooxygenase gene by plant-mediated RNAi impairs larval tolerance of gossypol. Nat Biotechnol 25: 1307-1313 (2007).
- Percival AE, Wendel JF, Stewart JM: Taxonomy and germplasm resources. In: Smith CW, Cothren JT (eds) Cotton: Origin, History, Technology, and Production, pp. 33-63. John Wiley & Sons, Inc., New York (1999).
- Smith CW, Cothren JT: Cotton: Origin, History, Technology, and Production. John Wiley & Sons, Inc., New York (1999).
- Sunilkumar G, Campbell LM, Puckhaber L, Stipanovic RD, Rathore KS: Engineering cottonseed for use in human nutrition by tissue-specific reduction of toxic gossypol. Proc Natl Acad Sci U S A 103: 18054-9 (2006).
- Wendel JF, Cronn RC: Polyploidy and the evolutionary history of cotton. Advances in Agronomy 78: 139-186 (2003).